 
Taylor ported the game Doom to Linux, as well as many other systems, during his spare time. The beginning of Linux as a gaming platform for commercial video games is widely credited to have begun in 1994 when Dave D. 1994–1997 Doom was one of the first major commercial games to be released for Linux. As the operating system itself grew and expanded, the amount of free and open-source games also increased in scale and complexity, with both clones of historically popular releases beginning with BZFlag, LinCity, and FreeCiv, as well as original creations such as Rocks'n'Diamonds and Tux Racer. 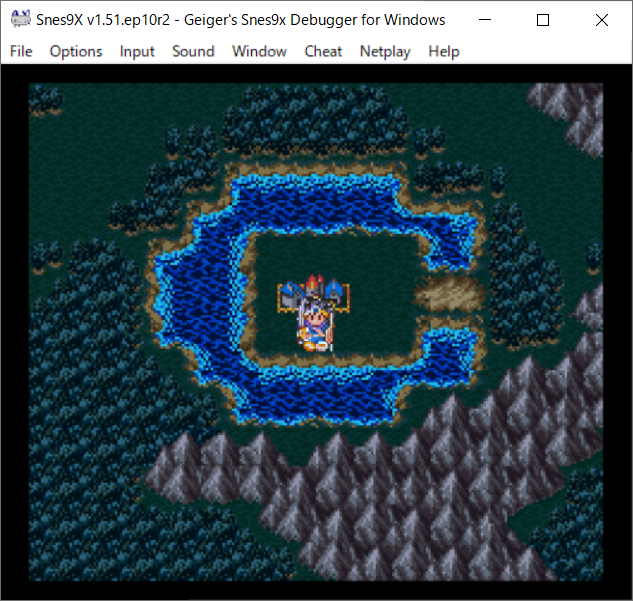
Popular early titles included Netrek and the various XAsteroids, XBattle, XBill, XBoing, X-Bomber, XConq, XDigger, XEmeraldia, XEvil, XGalaga, XGammon, XLander, XLife, XMahjong, XMine, XSoldier, XPilot, XRobots, XRubiks, XShogi, XScavenger, XTris, XTron, XTic and XTux games using the X Window System. The free software philosophy and open source methodology which drove the development of the operating system in general also spawned the creation of various early free games. A notable example of this are the " BSD Games", a collection of interactive fiction and other text-mode amusements. These games were mostly either arcade and parlour type games or text adventures using libraries like curses. Linux gaming started largely as an extension of the already present Unix gaming scene, which dates back to that system's conception in 1969 with the game Space Travel and the first edition in 1971, with both systems sharing many similar titles. Multi-drug resistance was also observed in eight species.See also: Open source video game § History NetHack, a primordial Unix game Several isolates exhibited no zone of inhibition, indicating they were resistant to the maximum concentration of the assay. Minimum inhibitory concentrations were determined for bacteria resistant to azithromycin, ciprofloxacin, and tetracycline. Resistant species included common isolates from the environment and pathogens of humans and fish. From these isolates, 39 bacteria species were identified by partial sequencing of the 16S ribosomal RNA gene. These samples were examined for resistance to six compounds representing major classes of antimicrobials and resistance was observed in 94 isolates. To improve the understanding of antimicrobial resistance in two river systems in Kansas, intestinal contents from 20 Channel Catfish (Ictalurus punctatus) and water samples were taken at eight sites on the Arkansas and South Fork Ninnescah rivers during the spring of 2012. 
Due to these potential threats, antimicrobial resistance in the aquatic environment should be closely monitored. Subsequently, resistance genes become established in natural systems and pose threats to human health and ecological processes. When antimicrobial-resistant bacteria enter the aquatic environment, water acts as a physical pathway for their distribution. Overuse of these compounds in clinical and agricultural applications has led to rapid evolution and global spread of antimicrobial resistance and rivers are the main receiving body for antimicrobials and resistant bacteria from urban effluents and agricultural runoff. Antimicrobial compounds have been used by humans to counteract bacterial infections since 1910. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
